 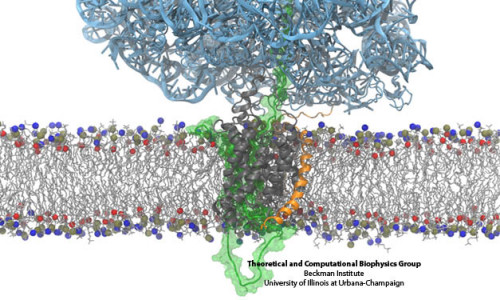
* Repeat the steps with the other molecule. Color by ID (red), Select again drawing method Isosurface with an Isovalue of -3. * Create a new representation, Color by ID (blue), Select drawing method Isosurface. * In Graphics Representation, change the standard representation to New Cartoon. Select the corresponding dx grid file from your apbs output Press Load. In the respective pqr files, what are the net charges for the two proteins _?_ * To visualize the electrostatic potential, select in the vmd main window the molecule 1yer. WHICH FILE FORMATS CAN BE IMPORTED AND EXPORTED BY NNDTOOLS imports: pmx, pmd, vmd, vpd - Yes exports: pmx, vmd, vpd - Yes imports, exports mqo - No exports pmd - No Ability to import. * Repeat for both structures and unpack the *.dx.gz file. MMDTOOLS MODEL JAPANESE TO ENGLISH TRANSLATION FEATURES 1. You might want to look at the program output and error logs if there are errors or warnings. 
* Again, save the output files, espcially the *.dx.gz grid files. * Save the output files (xxxx_aligned.in, xxxx_aligned.propka and xxxx_aligned.pqr) on disk. * Upload one of the aligned coordinate files.Ĭhoose the force field (AMBER) and naming scheme (AMBER), and click ’Submit’. zip, 600k, individual files) VMD can be used to load files that contain quantum mechanics (QM) data such as GAMESS log files and Molden files. VMD Quantum Chemistry Visualization Tutorial ( pdf, 2.0M) (required tutorial files. The proteins should show up as aligned entities. Tutorial works on Windows, Mac, and Unix/Linux platforms. This nearly replicates Qutemol-like rendering, but you’ll want to play with the colors that you use.Goal: Visualize possible interaction sites on proteins * Start VMD again and use the aligned Proteins from the previous section (load the previously saved coordinates). 
To give a soft appearance that emphasizes depth, we can use Tachyon with a diffuse material. Now, the result of our rendering can differ greatly depending on which renderer we use. As well, we assume you are a scientist and read the appropriate literature. We encourage users to refer to the Amber Reference Manual for syntax and detailed explanations. As for the rest of the settings, see the previous tutorial for turning on ambient occlusion, setting the material to diffuse, and Drawing Method as CPK with Sphere Scale/Bond Scale 1.6 and 1.0. Amber Tutorials Below are tutorials prepared by the Amber developers to help you learn how to use the Amber software suite. You can choose an alternative option by going to Graphics>Colors and choosing the Color Scale tab (I’ll actually be using red/green/blue today with an offset of 0.30 to make the colors lighter). The color gradient by default is red/white/blue. Now we can go back to the Draw style tab and choose coloring method as Timestep. Since there are 15 frames numbered from 0 to 14 and we want to see all of them, we’ll write 0:14 in the Draw Multiple Frames dialog box. Go to Graphics>Representations and select the Trajectory tab. In VMD, we can also overlay all of the different structures from our trajectory at the same time. Go to Extensions>Analysis>RMSD Trajectory Tool and you’ll see a window that looks like this: 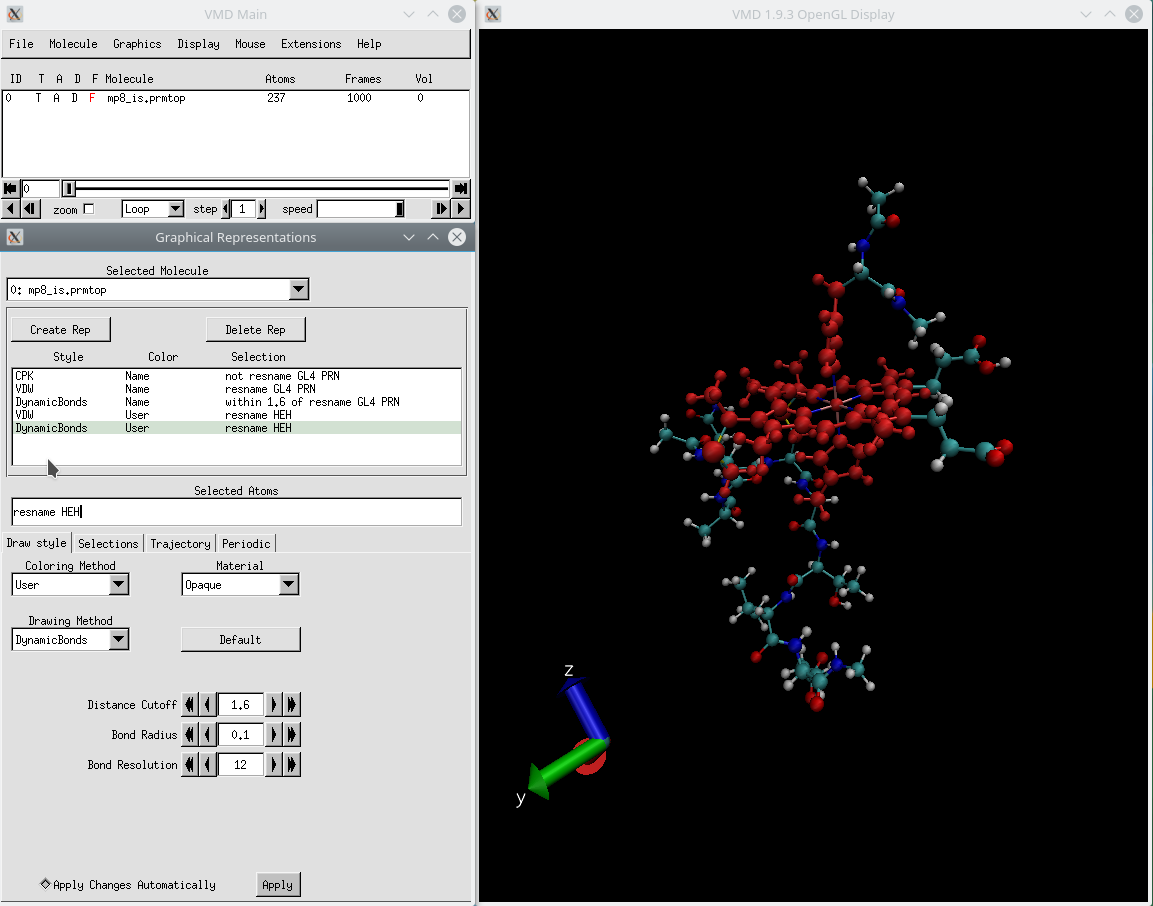
You’ll notice that the different structures are not aligned in any way. You can look at the different coordinates by sliding back and forth the bar on the bottom of the VMD main window. This tutorial is part of a detailed guide on making an abstract scientific. When you open this file, you’ll notice that there is more than one set of coordinates in it. Use VMD to create nice graphical representations of a large molecule. Start by opening the PPant-Anim.xyz file in VMD. Now, there are a few details of VMD we didn’t get to last time that are useful for getting more descriptive figures from your coordinate files with VMD. Revisiting VMD: Trajectories, Tachyon and beyond. Nevertheless, all you really need to know is that it is the example molecule we’ll work with today, and you can download the coordinates we’ll use for the visualization here and here.ġ. This prosthetic tether forms a thioester linkage to amino acids and delivers them to the active sites of many different enzymes found most commonly in natural product biosynthetic pathways. We’ll use a model for the phosphopantetheine tether of acyl carrier proteins. As a continuation of the previous tutorial, I will show you a few more things you can do with VMD along with some tricks for PyMOL. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
